09. Causal Effects and Adjusted Marginal Contrasts (RMST)
Source:vignettes/causal-rmst.Rmd
causal-rmst.RmdMoving Beyond the Hazard Ratio
In clinical trials and observational studies, researchers often wish to compare survival outcomes between two groups. Historically, this is answered using the Hazard Ratio (HR) from a Cox Proportional Hazards model. However, the HR is non-collapsible—meaning the omission of unmeasured covariates will mathematically bias the effect toward the null—and strictly relies on the proportional hazards assumption. If survival curves cross, the HR becomes mathematically invalid.
SuperSurv solves this by evaluating group differences on
the absolute time scale using the Restricted Mean Survival Time
(RMST) via G-computation (Standardization) on top of our
Ensemble Super Learner.
RMST calculates the area under the survival curve up to a specific time horizon, . By comparing the expected RMST if everyone in the dataset belonged to Group 1 versus if everyone belonged to Group 0, we obtain a robust, absolute measure of the difference:
Philosophy: “Causal Effect” vs. “Marginal Contrast”
How you interpret this depends entirely on the nature of your exposure variable. The math of G-computation is identical for both, but the statistical terminology must be used responsibly.
-
Causal Average Treatment Effect (ATE): You can
claim a Causal Effect if your variable is a manipulable
intervention. Examples include administering a drug, performing
a surgery, or applying a policy.
- Interpretation: “Administering this drug causally adds an average of 4.2 months of life over a 5-year period compared to the placebo.”
-
Adjusted Marginal Contrast: You must claim an
Adjusted Marginal Contrast if your variable is an
immutable trait or biological group. Examples include
biological sex, race, or a genetic biomarker. Because you cannot
“causally” intervene to change someone’s genetics, we are simply
comparing two groups while rigorously adjusting for all other
confounding variables.
- Interpretation: “After adjusting for all baseline clinical covariates, the presence of this biomarker is marginally associated with 4.2 additional months of survival over a 5-year period.”
Estimating the Effect with SuperSurv
Let’s demonstrate this using the built-in metabric
dataset. We will evaluate the effect of the binary biomarker
x4 (1 = present, 0 = absent). Because
x4 is a biomarker, we will interpret the result as an
Adjusted Marginal Contrast.
library(SuperSurv)
set.seed(123)
# Load built-in data
data("metabric", package = "SuperSurv")
# Define predictors and time grid
X <- metabric[, grep("^x", names(metabric))]
new.times <- seq(10, 150, by = 10)1. Train the Super Learner
First, train the ensemble. We must set
control = list(saveFitLibrary = TRUE) so the models are
saved for the G-computation prediction phase.
2. Estimate the Adjusted Marginal RMST Contrast
We use the estimate_marginal_rmst() function to compute
an adjusted marginal contrast on the RMST scale. The function sets the
binary grouping variable x4 to 1 for all patients, predicts
their survival curves, integrates those predictions up to the
restriction horizon tau, and then repeats the same
procedure with x4 set to 0. The difference between these
two standardized averages yields the adjusted RMST contrast.
# Estimate the adjusted difference up to tau = 100 months
results <- estimate_marginal_rmst(
fit = fit,
data = metabric,
trt_col = "x4",
times = new.times,
tau = 100
)
#> Adjusted Delta RMST at tau = 100: -1.27 time units
print(results$ATE_RMST)
#> [1] -1.269855Interpretation: If the resulting
RMST
value is -1.24, this indicates that, after standardizing
over the observed covariate distribution using the fitted Super Learner
ensemble, the group with x4 = 1 is predicted to have
approximately 1.24 fewer months of restricted mean survival than the
group with x4 = 0 over a 100-month horizon.
Uncertainty: To quantify uncertainty,
estimate_marginal_rmst() can optionally apply a
perturbation-based inference procedure conditional on the fitted
ensemble. This returns a perturbation-based standard error, confidence
interval, and Wald-type p-value.
rmst_results_inf <- estimate_marginal_rmst(
fit = fit,
data = metabric,
trt_col = "x4",
times = new.times,
tau = 100,
inference = TRUE,
B = 100,
seed = 123
)
#> Adjusted Delta RMST at tau = 100: -1.27 time units | SE = 0.026 | 95% CI = [-1.32, -1.219]
rmst_results_inf$ATE_RMST
#> [1] -1.269855
rmst_results_inf$SE_RMST
#> [1] 0.02571039
rmst_results_inf$CI_RMST
#> lower upper
#> -1.320246 -1.219464
format.pval(rmst_results_inf$p_value, digits = 3, eps = 1e-16)
#> [1] "<1e-16"Note: Because this perturbation procedure conditions on the final fitted SuperSurv model and does not refit the learner library or ensemble weights, the resulting confidence interval reflects conditional uncertainty for the standardized RMST contrast and may be relatively narrow.
3. Visualizing the Effect Over Time
The difference between groups might be near zero early on but
substantial later. We can visualize how the adjusted RMST contrast
evolves across different restriction times using
plot_marginal_rmst_curve(). When
inference = TRUE, the function also displays
perturbation-based confidence intervals as a ribbon.
# Plot the Delta RMST across a sequence of tau values
tau_grid <- seq(20, 140, by = 30)
plot_marginal_rmst_curve(
fit = fit,
data = metabric,
trt_col = "x4",
times = new.times,
tau_seq = tau_grid,
inference = TRUE,
B = 100,
seed = 123,
ci_level = 0.95
)
#> Adjusted Delta RMST at tau = 20: -0.01 time units | SE = 0 | 95% CI = [-0.01, -0.009]
#> Adjusted Delta RMST at tau = 50: -0.204 time units | SE = 0.005 | 95% CI = [-0.214, -0.194]
#> Adjusted Delta RMST at tau = 80: -0.781 time units | SE = 0.014 | 95% CI = [-0.807, -0.754]
#> Adjusted Delta RMST at tau = 110: -1.567 time units | SE = 0.027 | 95% CI = [-1.619, -1.514]
#> Adjusted Delta RMST at tau = 140: -2.649 time units | SE = 0.045 | 95% CI = [-2.737, -2.56]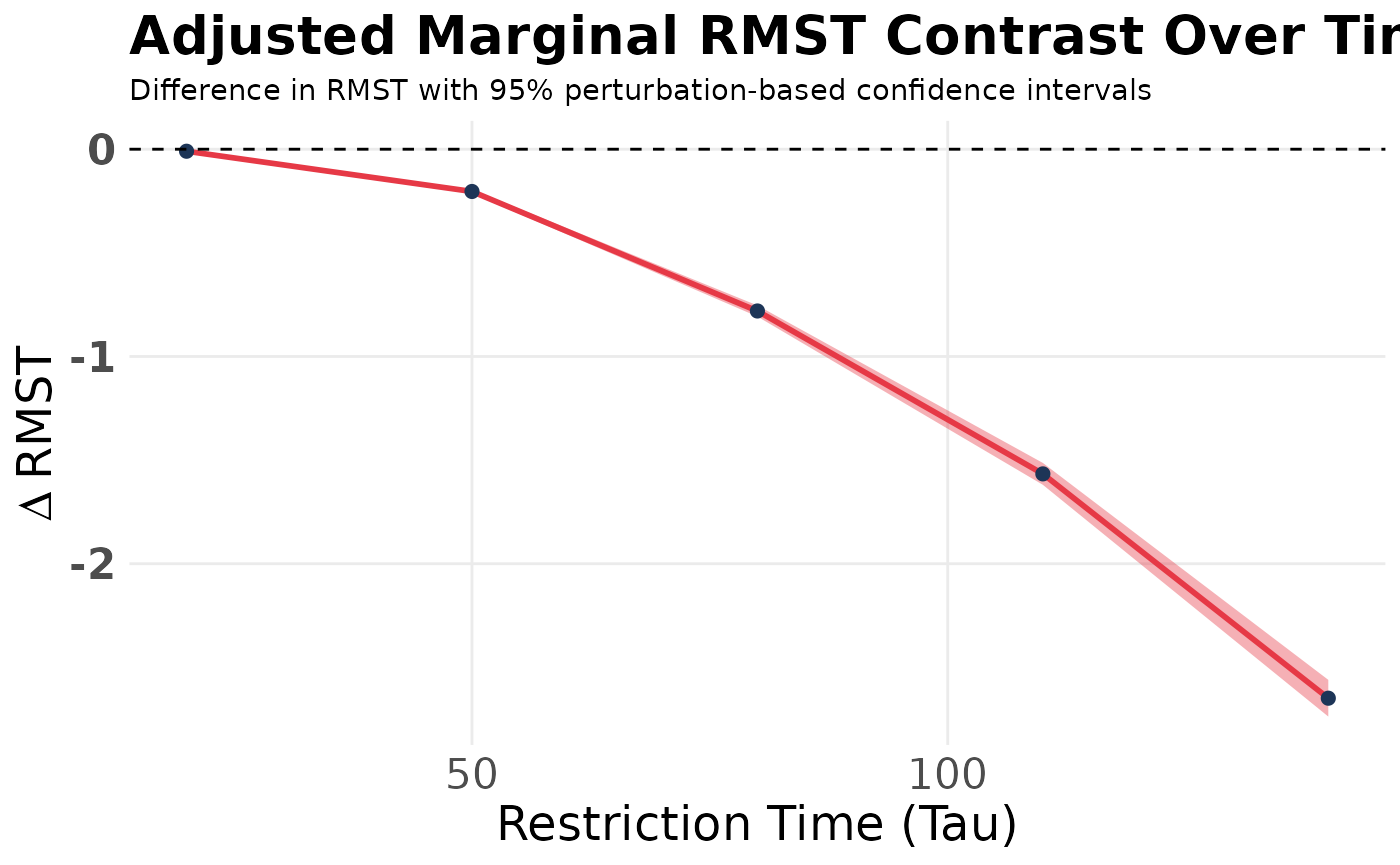
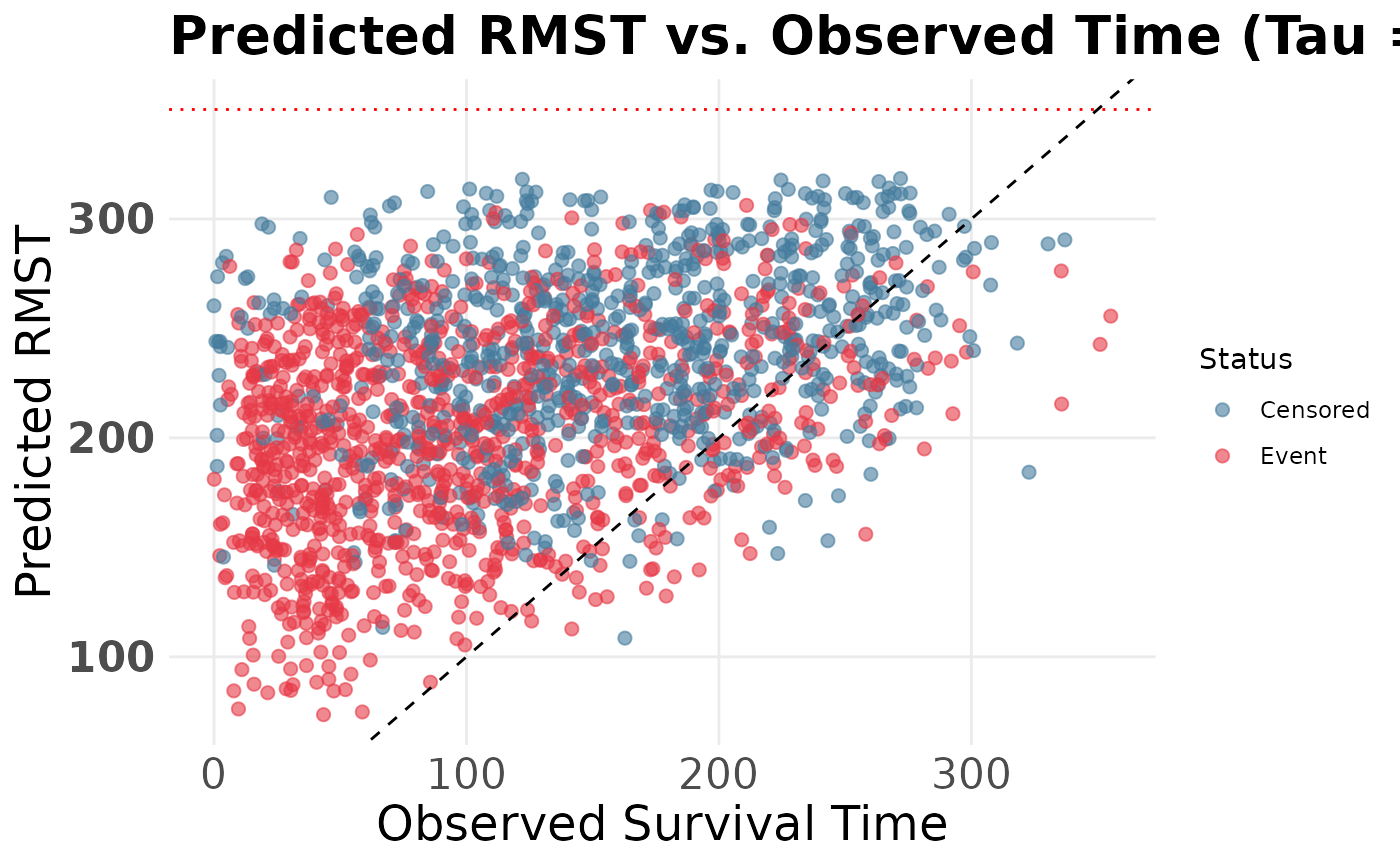
4. Diagnostic: Predicted RMST vs. Observed Time
To evaluate how well our model’s restricted expectations align with reality, we can plot the predicted RMST for the observed data against their true survival times. Patients who experienced the event should lie close to the diagonal line up to .
plot_rmst_vs_obs(
fit = fit,
data = metabric,
time_col = "duration",
event_col = "event",
times = new.times,
tau = 350
)